Touchdown pcr faq12/22/2023  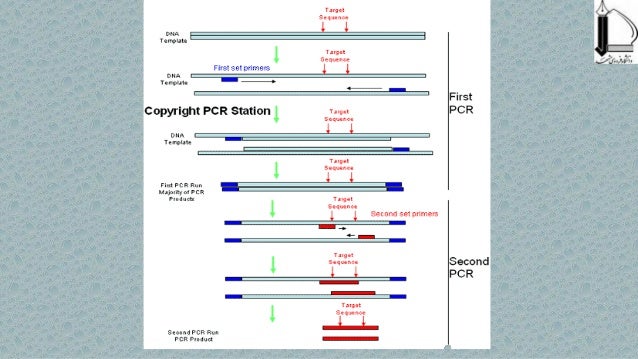
For calculating the T m values by nearest-neighbor thermodynamic models, one of the following calculators is recommended: See Troubleshooting section for information about how various PCR conditions and additives affect melting temperature. The latter is used more frequently because the calculations are simple and can be done quickly by hand. The former will give more accurate T m estimation because it takes into account the stacking energy of neighboring base pairs. However, nearest-neighbor thermodynamic models are preferred over the more conventional calculation: T m ≈ 4(G-C) + 2(A-T). Although there are several T m calculators available, it is important to note that these calculations are an estimate of the actual T m due to lack of specific information about a particular reaction and assumptions made in the algorithms for the T m calculators themselves. Knowing the melting temperature (T m) of the primers is imperative for a successful PCR experiment. There are a few common problems that arise when designing primers: 1) self-annealing of primers resulting in formation of secondary structures such as hairpin loops ( Figure 1a) 2) primer annealing to each other, rather then the DNA template, creating primer dimers ( Figure 1b) 3) drastically different melting temperatures (T m) for each primer, making it difficult to select an annealing temperature that will allow both primers to efficiently bind to their target sequence during themal cycling ( Figure 1c) (See the sections CALCULATING MELTING TEMPERATURE (T m) and MODIFICATIONS TO CYCLING CONDITIONS for more information on T ms). When designing a set of primers to a specific region of DNA desired for amplification, one primer should anneal to the plus strand, which by convention is oriented in the 5' → 3' direction (also known as the sense or nontemplate strand) and the other primer should complement the minus strand, which is oriented in the 3' → 5' direction (antisense or template strand). Troubleshoot failed PCR experimentsĭesigning appropriate primers is essential to the successful outcome of a PCR experiment.Design and optimize a PCR experiment for any DNA template.Understand the function of various reaction components and their overall effect on a PCR experiment. 
Set up reactions and thermal cycling conditions for a conventional PCR experiment.By following this PCR guide, students should be able to: This protocol outlines the basic principles of PCR, provides a methodology that will result in amplification of most target sequences, and presents strategies for optimizing a reaction. PCR failures can become frustrating unless patience and careful troubleshooting are employed to sort out and solve the problem(s). Another potential problem occurs when mutations are unintentionally introduced in the amplicons, resulting in a heterogeneous population of PCR products. When PCR fails it can lead to many non-specific DNA products of varying sizes that appear as a ladder or smear of bands on agarose gels. While straightforward and generally trouble-free, there are pitfalls that complicate the reaction producing spurious results. PCR is a powerful amplification technique that can generate an ample supply of a specific segment of DNA (i.e., an amplicon) from only a small amount of starting material (i.e., DNA template or target sequence). Automation and refinement of this technique progressed with the introduction of a thermal stable DNA polymerase from the bacterium Thermus aquaticus, consequently the name Taq DNA polymerase. The theoretical process was outlined by Keppe and coworkers in 1971 however, it was another 14 years until the complete PCR procedure was described and experimentally applied by Kary Mullis while at Cetus Corporation in 1985. The development of the polymerase chain reaction (PCR) is one of those innovations that changed the course of molecular science with its impact spanning countless subdisciplines in biology. For example, the field of microbiology was transformed with the advent of Anton van Leeuwenhoek's microscope, which allowed scientists to visualize prokaryotes for the first time. In the biological sciences there have been technological advances that catapult the discipline into golden ages of discovery. 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
